Biorepository Development
Development of Biorepository FFPE and Plasma Database and UI based on clinically relevant Oncomine variants
Bioninformatician | Below Average Basketball Player | Lover of Code and Science
Welcome on in...
Innovative and results-driven Bioinformatician with a passion for transforming complex biological data into actionable insights. With a strong foundation in computational biology, data science, and software development, I aim to contribute to pioneering research in genomics and precision medicine.
Development of Biorepository FFPE and Plasma Database and UI based on clinically relevant Oncomine variants
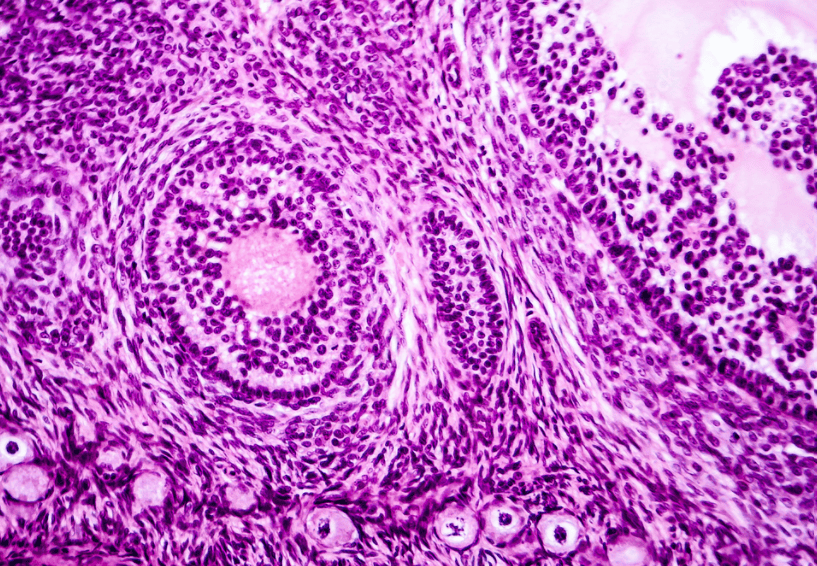
Inception V3 & ViT Model Training, Validation, and Testing for Deep Learning Classification of genetic variant predictions in H&E Images

Supervised and Neural Network Machine Learning Model development to classify SNV/INDEL Ion Torrent variations
Develop R&D Bioinformatics App UI to assist colleagues in analyzing data, connecting internal databases, and running approved 3rd-party software
Professional/Educational journey and key accomplishments
2008
Specialized in genomic data analysis and machine learning applications in healthcare. Thesis focused on novel algorithms for variant calling in cancer genomics.
March 2011
Published "Novel Machine Learning Approaches for FFPE Sample Analysis" detailing new computational methods for working with challenging sample types.
2012 - 2015
Developed pipelines for processing next-generation sequencing data. Implemented quality control measures that improved data reliability by 15%.
2014 - 2016
Specialized in genomic data analysis and machine learning applications in healthcare. Thesis focused on novel algorithms for variant calling in cancer genomics.
2016 - 2019
Contributed to the development of analysis tools for Ion Torrent sequencing platform. Collaborated with cross-functional teams to implement new features for variant detection.
2019 - 2020
Contributed to the development of analysis tools for Ion Torrent sequencing platform. Collaborated with cross-functional teams to implement new features for variant detection.
2020 - 2023 (Sr. Scientist)
2023 - Current (Staff Scientist)
Contributed to the development of analysis tools for Ion Torrent sequencing platform. Collaborated with cross-functional teams to implement new features for variant detection.
2023 - 2024
Spearheaded development of deep learning models for histopathology image analysis, achieving 92% accuracy in predicting genetic variants from H&E slides.
Jan 2026
Analytical and Clinical Validation of the IVD-Compliant Oncomine Dx Express Test for Genomic Profiling of FFPE Tumor Samples
(Hover and Click on Dots/Lines to get more information)